Blog | Reading Time 6 minutes
OMICS research can help answering shrimp farmers practical questions
Studying and managing shrimp gut and water microbiota
The main microbial components of shrimp farming systems are the water, the soil, and the shrimp gut microbiota. Lallemand Animal Nutrition has developed in-feed probiotics and bioremediation solutions that can help target the modulation of gut and water microbiota, respectively. In turn, it helps secure shrimp farming outcomes. What is their real impact on these microbial ecosystems? Thanks to metagenomics, recent studies conducted by the Lallemand Monogastric Center of Excellence help us to answer some practical questions often asked by shrimp farmers in hatcheries and nurseries.
Trial setup
We carried out a controlled experiment in Vietnam covering both the hatchery and nursery phases of white leg shrimp (Litopenaeus vannamei). The trial evaluated the use of a water bioremediation product (LALSEA PL Pack, Lallemand Animal Nutrition) alongside an in-feed probiotic (BACTOCELL or LALPACK PROBIO, Lallemand Animal Nutrition) in comparison to, at the hatchery phase, water antibiotic prophylaxis and, at the nursery phase, competitor’s microbial solutions or negative control (Figure 1).
[table id=14 /]
Using amplicon sequencing1 technique as a metagenomic approach, it was possible to assess the changes in gut and water microbiota induced by the different microbial solutions and the consequences on performance. Here, the application of state-of-the-art genomic techniques enabled researchers to answer practical questions related to positive microbiota management in shrimp farming:
- Does the water microbiome impact all life stages of shrimp similarly? No.
In the hatchery phase, the gut and water microbial populations were very similar, unlike in the nursery phase. In the later stages, shrimp and water microbiota progressively differentiate as the gut microbiota becomes much more influenced by the feed and by its host, the shrimp. This suggests that, at later stages, the use of both specific in-feed probiotic and in-water (bioremediation) bacteria will be necessary to target these well-differentiated microbial compartments.
- Can we track the added beneficial bacteria in the water and in the shrimp? Yes.
The bacteria from the water bioremediation solution (Bacillus and Pediococcus) were tracked back to the genus level. Both could be found in the water. Similarly, at the nursery phase, the bacteria from the probiotic solution could be found in the shrimp gut (Figure 2).
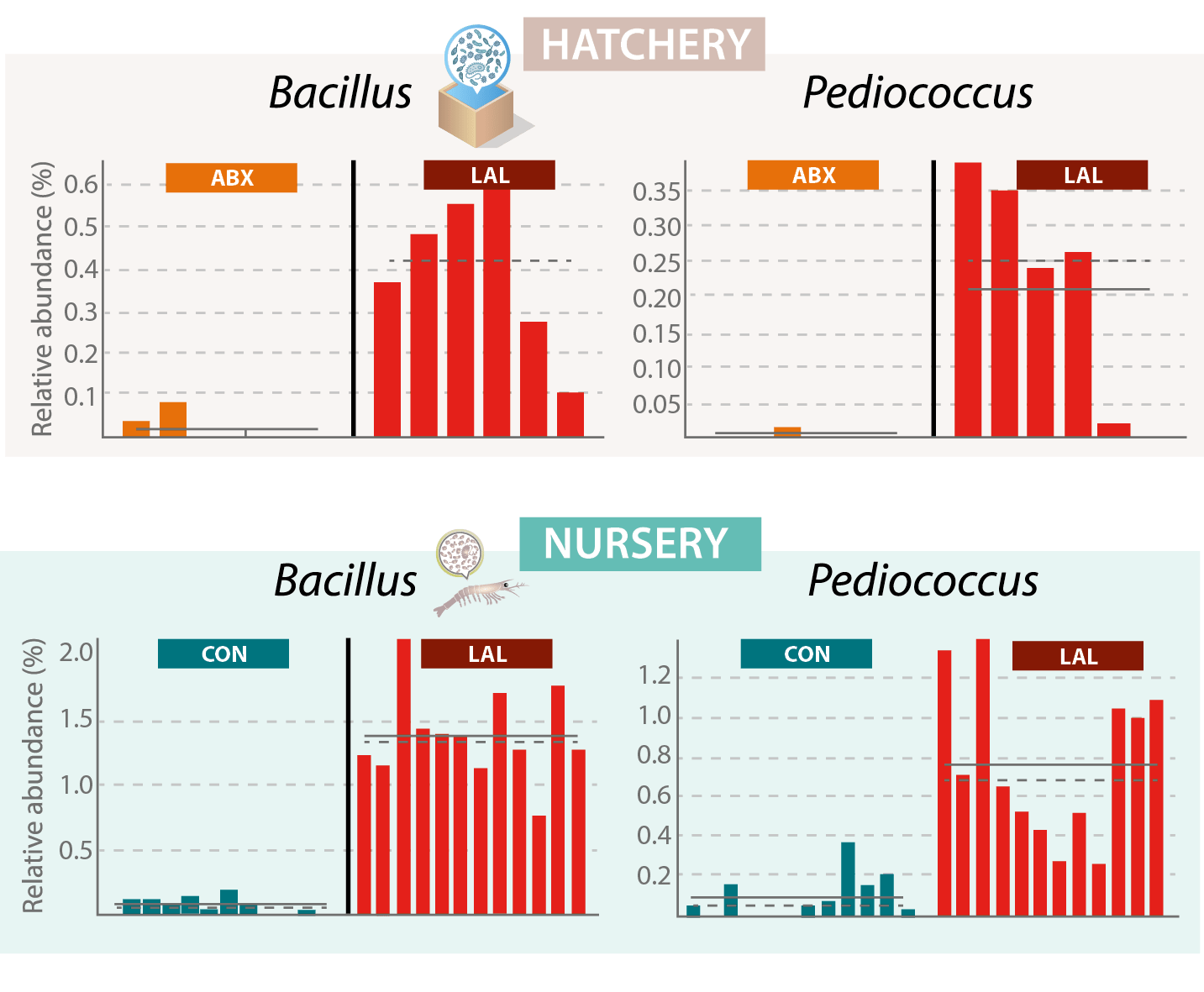
Figure 2. Relative abundance of Bacillus and Pediococcus in ABX and LAL treatments in hatchery water and nursery shrimp gut microbiota (P<0.05); the individual data, average (straight line), and median (dotted line) for each treatment are shown.
- Do probiotic and bioremediation products reduce the prevalence of undesirable bacteria? Yes.
In the hatchery, a number of undesirable bacterial groups detected in the water of the antibiotic group were not identified when using our products, including a family of Vibrio species and a genus linked to early mortality syndrome transfer in shrimp.
Similarly, in the nursery phase, the application of our solutions significantly reduced the prevalence of potential pathogens in the shrimp gut from 7.5% to 0.28%, including Vibrio, Acinetobacter, Thalassomonas, Owenweeksia, and Sedimenicola. The prevalence of the Enterobacteriaceae family was reduced; there was no Cellvibrio, which hydrolyses chitin; and no Coccinistipes were detected. This last genus is typically favored by some microalgal bloom and generally found in locations relatively rich in organic carbon.
- Can we see effects on the presence of beneficial bacteria? Yes.
In both stages, researchers could see enrichment of shrimp gut microbiota with beneficial bacteria.
These include Thalassobious from the Rhodobacteriaceae family at the nursery phase. This family has been recently suggested as a source of probiotics as it is able to synthesize vitamin B12 and tropodithietic acid (antagonistic to Vibrio species) while having a large potential contribution to feed digestibility. A higher proportion of Psychroserpens was also found, in line with previous findings after probiotic feeding.
In the nursery phase, researchers noticed an increase in Phaeobacter, a genus affiliated with the Rhodobacteriaceae family, and proposed as probiotic for its involvement in the biosynthesis of natural antimicrobial compounds and nitrate reduction.
In the water, Bacillus were detected only in the group with Lallemand products, together with a significantly higher proportion of Sphingomonas, which are involved in denitrification; Rubrivivax, which carry nitrogen and help with carbon fixation; and Kaistobacter, which have the capacity to degrade toxic aromatic compounds.
Maintaining a positive microbiota both in the shrimp gut and the water has several benefits such as: support for the immune system and animal robustness; improved nutritional performance and water quality. This helps contribute to lower pathogen pressure, which, in turn, reduces the likelihood of disease outbreaks. Together, our findings support existing knowledge on the capacity of specific microbial solutions, both in-feed and in water, to reinforce the presence of beneficial microorganisms while helping to keep potential pathogens at bay.
- Are all probiotic and bioremediation products acting the same way? No.
When comparing Lallemand products and the alternative products, it appears that the Lallemand solutions were more effective at maintaining the shrimp gut with beneficial microbial groups, and, as a result, at reducing the relative abundance of undesirable bacteria. This emphasizes the importance of strain specificity in microbial based products.
- Finally, does this translate into performance? Yes.
End-point body weight, feed conversion ratio (FCR) and survival were improved in the LAL group compared to the COMP group in the nursery phase as a result of good water quality and balanced intestinal microbiota. The LAL group yielded better and more consistent performance.
Conclusion
The latest molecular tools are effective to shed some light in the understanding of microbiota associated with shrimp farming and their positive manipulation by microbial solutions (probiotics and bioremediation products). The microbial inputs can be tracked in the water and in the shrimp gut. They can help reduce undesirable bacteria pressure and increase the presence of symbionts.
The choice of a product containing carefully selected and well-documented bacterial strains is crucial to deliver the expected effects of microbial solutions at the farm level — promoting a positive microbial environment in and around the shrimp that secures healthy stock and robust growth.
For more results, download our white paper!
Published May 11, 2021 | Updated May 29, 2023
Related articles
Need specific information?
Talk to an expert


